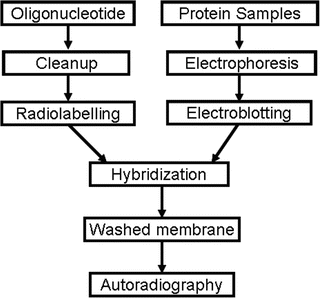
The technique usually requires either heat inactivation (2) or partial purification (3-5) of crude protein extract. One of these filters is used to visualize proteins in the lanes by staining, and the other one is for detecting protein factors interacting with a labeled probe (1). The polyacrylamide gel containing the proteins is then sandwiched between two filters so that the proteins can diffuse out and bind onto the filters to produce replicas of the gel. (1) In this original protocol, proteins are first separated based on size by SDS-PAGE. The technique is modified from a previous procedure called 'protein blotting' described by Bowen et al. Southwestern blotting is a technique developed for the detection and characterization of protein factors interacting with distinct DNA motifs. Identification of protein factors that bind to genes to turn them on or off is therefore important in investigating gene functions. 
Transcription and hence expression regulation of a gene involves initially highly specific interactions of protein factors with distinct short DNA sequence motifs, 6-8 bp in size, usually located within the promoter or other functional cis-acting elements. Southwestern blotting is useful in investigating DNA-protein interactions to aid the identification of DNA-binding protein factors. Understanding the spatial and temporal expression patterns of a gene is essential in elucidating its functions. In this article, a commonly used protocol and expected results are described and discussed. The whole Southwestern blotting procedure takes ~4 d to complete. This technique could be combined with database searching (TransFac, ), prediction of potential protein factors binding onto a target motif (e.g., Patch search), in vitro supershift EMSA and in vivo chromatin immunoprecipitation (ChIP) assays for effective identification of protein factors. The interaction of the probe with the protein(s) is later visualized by autoradiography. This method combines the features of Southern and Western blotting techniques a denaturing SDS-PAGE is first employed to separate proteins electrophoretically based on size, and after transferring the proteins to a membrane support, the membrane-bound proteins are renatured and incubated with a P-labeled double-stranded oligonucleotide probe of specific DNA sequence. The advantage of this technique over other related methods such as electrophoretic mobility shift assay (EMSA) and DNA footprinting is that it provides information regarding the molecular weight of unknown protein factor. Southwestern blotting is used to investigate DNA-protein interactions.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
